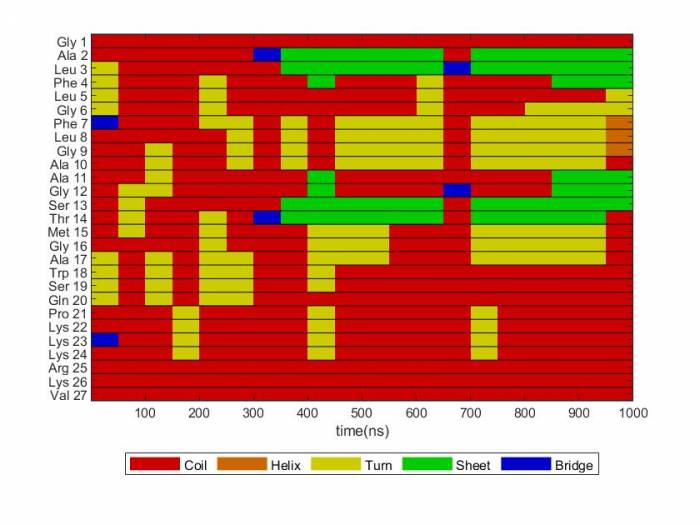
Secondary Structure using DSSP
The code is adopted and changed from the following script for secondary structure calculation in VMD. https://www.ks.uiuc.edu/Research/vmd/script_library/scripts/sscache/
This analysis computes the secondary structure of protein residues in time for a dcd trajectory. The scripts needed can be downloaded from below.
Steps
- change the psf and dcd in ss_content.tcl according to where these files are in your directory
- change the last_file number in ss_content.csh according to the number of trajectories. If you are using a supercomputer other than deepthought2 change this file accordingly.
- run ss_content.csh ( sbatch ss_content.csh)
- this will give you multiple .dat files containing the secondary structure in each trajectory. you can combine these into a single secondary_structure file for further analysis.This can be done by the following code:
for i in {1..40};do
sed '1~10!d' sec_structure-$i.dat >> ss_content.dat
done
the line sed '1~10!d' in above means, we are only taking every 10th line of the secondary structure vs time.
To visualize the secondary structure you can use the Matlab file ss_content.m you need to change a few things in this file: num_residues, num_timesteps, step-size, and residue names at the end. You can run this file locally on you pc/mac or on the cluster